Genetic and Phylogenetic Analysis of Skin Mites in Wild and Domestic Mammals
In a study recently published in PLoS ONE, researchers reported the presence, prevalence, and phylogenetic relationships of skin mites, including Demodex mites, in social wild and domestic mammals.
In a study recently published in
The subclass Acari contains seven orders, one of which is
DNA sequence analysis has proven useful in establishing phylogenetic relationships within the Demodecidae family. The mitochondrial
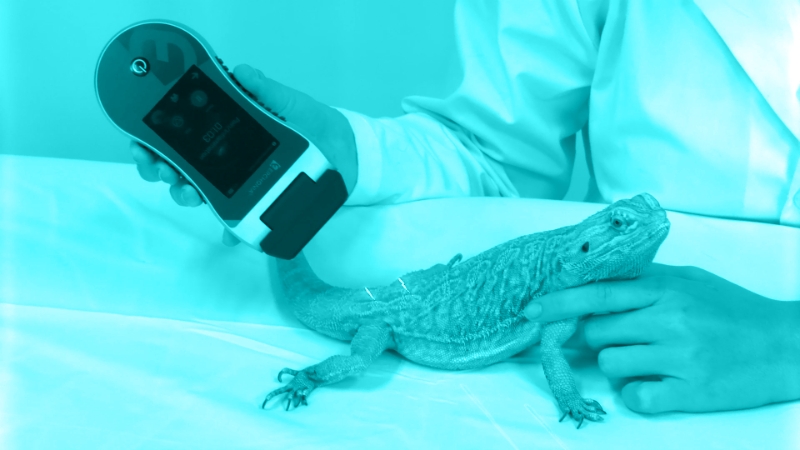
Authors collected hair samples from healthy gray wolves (Canis lupus), yellow-bellied marmots (Marmota flaviventris), and four species of insectivorous bats (Myotis californicus, M.lucifugus, M. yumanenses, Corynorhinus townsendii); the wolves were captive, and the marmots and bats were free-ranging. These animals were chosen because of their
Hair samples were also collected via skin scrapings from one of the authors (who was healthy), and domestic dogs, cats, and a ferret with demodicosis. Several Demodex species were identified per host using microscopic and morphologic examination: human (D. folliculorum), dog (D. canis, D. injai), cat (D. gatoi, D. felis), and ferret (Demodex spp. similar to D. canis).
Authors used real-time quantitative PCR (qPCR) to amplify a fragment of the mitochondrial16S rRNA gene from the wild animal hair samples; 16S qPCR products meeting certain PCR parameters underwent DNA sequencing. Conventional PCR was used to amplify the16S rRNA gene in the human and domestic animal hair samples. The authors also used conventional PCR to amplify the18S rRNA gene, using samples sequenced successfully for the16S rRNA gene.
Authors searched
Real-time qPCR results positively identified mite DNA in 17% (5/28) of bat samples and 13% (2/16) of marmot samples. The presence of mite DNA in 14% (3/22) of wolf samples was not certain because of unsuccessful DNA sequencing in those samples.
16S rDNA sequencing demonstrated that the mites identified in bat and marmot hair samples were not Demodex mites, but rather, other mites within the Prostigmata order. Specifically, three different mite species were identified in three of the bat species; two mite species were identified in the one marmot species. Given these findings, authors suggested these mites could not “be assigned to the same genus based only on a common host, and seem[ed] to evolve according to the specific habitat and/or specific hair and sebaceous gland of the mammalian host.”
Authors searched the GenBank database for sequences matching the 16S rDNA and 18S rDNA sequences identified in hair samples from the bats, marmots, dogs, cats, ferret, and human. Several of the Demodex sequences identified in the dogs, ferret, and human matched previously-entered Demodex sequences from around the world (Austria, China, Spain, USA), indicating a global distribution of Demodex mites.
Authors determined the percentage of each nucleotide in the hair samples. Regarding Demodex mites, nucleotide diversity was markedly greater using the16S rRNA gene than the18s rRNA gene, supporting the previous
The wide distribution of Demodex species, particularly D. canis, raises questions on how D. canis has evolved, adapted, and been transmitted across the world and across species. Authors wrote that the “D. canis mite evolution may be a consequence of the relationship between ‘human-related’ mammals and humans.” In addition, they noted that studying mites in wolves could help determine whether mites evolved along with dog domestication.
Dr. JoAnna Pendergrass received her doctorate in veterinary medicine from the Virginia-Maryland College of Veterinary Medicine. Following veterinary school, she completed a postdoctoral fellowship at Emory University’s Yerkes National Primate Research Center. Dr. Pendergrass is the founder and owner of JPen Communications, LLC.