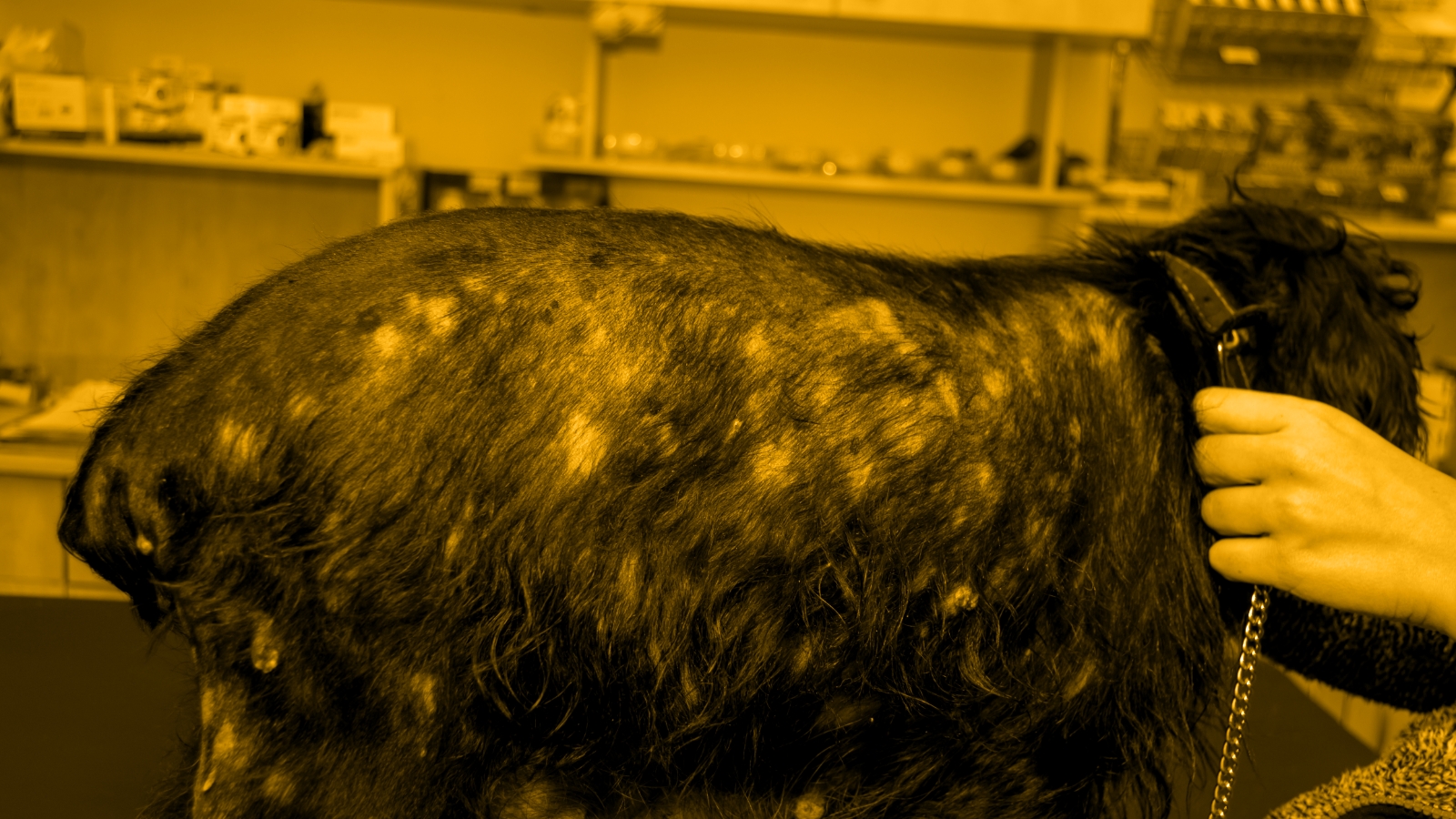
Bacterial Dysbiosis in Allergen-Induced Canine Atopic Dermatitis
A recent study reported bacterial dysbiosis in dogs sensitized to, then challenged with, house dust mite (HDM) allergens.
A recently published study in
Changes in the skin's microbiota occur in humans and dogs with atopic dermatitis (AD). Clinical studies in AD have focused on the Staphylococcus species of bacteria—
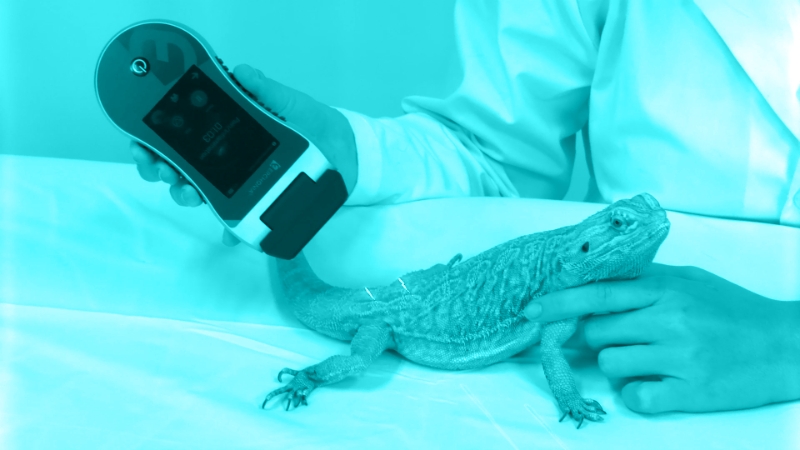
The study included eight atopic dogs previously sensitized with Dermatophagoides farinae (Df) HDM. The dogs were kept primarily indoors to prevent allergen exposure, but were allowed outside into their runs for exercise and interaction. For at least three months prior to skin sample collection, the dogs did not receive treatment with antibiotics, anti-inflammatory medications, or immunosuppressants.
To perform the allergen challenge, investigators painted 20 µL of a Df solution onto each dog’s right groin (challenged site) every 24 hours for 3 consecutive days; one dog received only two applications due to severe local lesions following the second application.
Investigators collected 12 skin samples from each dog by swabbing the right groin and left groin (nonchallenged site) at specific time points: pre-challenge, and days 1, 7, 14, 21, and 28 after the allergen challenge; they also swabbed the Df solution.
Investigators performed NGS of the bacterial 16S rRNA gene to evaluate temporal changes in the dogs’ skin microbiota. They also performed real time quantitative PCR (RT-qPCR) to quantify Staphylococcus spp. and S. pseudintermedius. In addition, investigators used indices of microbial diversity to evaluate the skin microbiota.
Prior to the allergen challenge and on day 1 after the challenge, investigators scored the dogs’ skin lesions (erythematous macules, edema, papules/pustules, excoriations) on a scale of zero (absent) to three (strong, severe).
All dogs developed skin lesions on the challenged site, but not on the unchallenged site. For each dog, skin lesion scores increased 24 hours after the final Df application. All skin lesions resolved on day 7 after the final application.
At any time point, no significant differences were observed in bacterial diversity (richness in bacterial species) between the challenged and unchallenged sites. Analyzing only the challenged site, the number of observed bacterial species was significantly higher in samples from day 21 than in pre-challenge samples.
Comparing the challenged and nonchallenged sites, pre-challenge samples did not demonstrate significant differences in bacterial microbiota, based on Kruskal-Wallis statistical testing and Linear discriminant analysis effect size. After allergen challenge, increases in Corynebacteriaceae (day 1) and Staphylococcaeae (days 7, 14, and 21) were observed on the challenged site; significant increases in the bacterial order Bacilli and class Bacillales, which contain the Streptococcaceae and Staphylococcaceae families, were observed on day 21. Relative proportions of the Staphylococcaeae family were not significantly different between the sites.
When analyzing only the challenged site, significant decreases in Fusobacteriaceae were noted on day 7 compared with day 1.
RT-qPCR results demonstrated no significant differences in Staphylococcus spp. and S. pseudintermedius copy numbers between pre-challenge and day 1 samples, indicating little change in these bacteria at the time of acute skin lesion development. Comparing the challenged site with the nonchallenged site, S. pseudintermedius copy numbers significantly increased on days 7 and 21; Staphylococcus spp. copy numbers significantly increased on day 21.
Df HDM microbiota analysis demonstrated an abundance of the Bartonellaceae and Leuconostocaceae families. Proportions of these bacterial families did not change in the skin samples, confirming the absence of bacterial contamination of the dogs’ skin by HDM allergens.
The nonsignificant changes in microbial diversity observed in this study may be due to the nature of the lesions (mild, localized only to the challenge area); it is also possible that changes occurred at time points different than those evaluated. Despite this, the significant increases in Staphylococcus spp. and S. pseudintermedius copy numbers indicate their importance in bacterial dysbiosis in allergen-induced CAD.
The authors noted the small sample size as a study limitation. They offered several suggestions for future longitudinal studies in dogs with allergen-induced or natural CAD, including analysis of microbial interactions during and after atopic skin lesion development.
Dr. JoAnna Pendergrass received her doctorate in veterinary medicine from the Virginia-Maryland College of Veterinary Medicine. Following veterinary school, she completed a postdoctoral fellowship at Emory University’s Yerkes National Primate Research Center. Dr. Pendergrass is the founder and owner of JPen Communications, LLC.